USP9X Deubiquitylates DVL2 to Regulate WNT Pathway Specification
- PMID: 31340145
- PMCID: PMC6884140
- DOI: 10.1016/j.celrep.2019.06.083
USP9X Deubiquitylates DVL2 to Regulate WNT Pathway Specification
Abstract
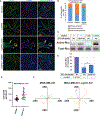
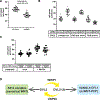
The WNT signaling network is comprised of multiple receptors that relay various input signals via distinct transduction pathways to execute multiple complex and context-specific output processes. Integrity of the WNT signaling network relies on proper specification between canonical and noncanonical pathways, which presents a regulatory challenge given that several signal transducing elements are shared between pathways. Here, we report that USP9X, a deubiquitylase, and WWP1, an E3 ubiquitin ligase, regulate a ubiquitin rheostat on DVL2, a WNT signaling protein. Our findings indicate that USP9X-mediated deubiquitylation of DVL2 is required for canonical WNT activation, while increased DVL2 ubiquitylation is associated with localization to actin-rich projections and activation of the planar cell polarity (PCP) pathway. We propose that a WWP1-USP9X axis regulates a ubiquitin rheostat on DVL2 that specifies its participation in either canonical WNT or WNT-PCP pathways. These findings have important implications for therapeutic targeting of USP9X in human cancer.
Keywords: DUB-E3 interactions; E3 ubiquitin ligases; USP9X; WNT signaling; WNT-PCP signaling; WWP1; deubiquitylases; non-canonical WNT signaling; ubiquitin rheostat.
Copyright © 2019 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors declare no competing financial interests.
Figures





References
-
- Allache R, Wang M, De Marco P, Merello E, Capra V, and Kibar Z (2015). Genetic studies of ANKRD6 as a molecular switch between Wnt signaling pathways in human neural tube defects. Birth Defects Res. A Clin. Mol. Teratol 103, 20–26. - PubMed
-
- Angers S, Thorpe CJ, Biechele TL, Goldenberg SJ, Zheng N, Mac-Coss MJ, and Moon RT (2006). The KLHL12-Cullin-3 ubiquitin ligase negatively regulates the Wnt-beta-catenin pathway by targeting Dishevelled for degradation. Nat. Cell Biol 8, 348–357. - PubMed
-
- Bernassola F, Karin M, Ciechanover A, and Melino G (2008). The HECT family of E3 ubiquitin ligases: multiple players in cancer development. Cancer Cell 14, 10–21. - PubMed
-
- Cadavid AL, Ginzel A, and Fischer JA (2000). The function of the Drosophila fat facets deubiquitinating enzyme in limiting photoreceptor cell number is intimately associated with endocytosis. Development 127, 1727–1736. - PubMed
-
- Chen HI, Einbond A, Kwak SJ, Linn H, Koepf E, Peterson S, Kelly JW, and Sudol M (1997). Characterization of the WW domain of human yes-associated protein and its polyproline-containing ligands. J. Biol. Chem 272, 17070–17077. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

