Protein identification in imaging mass spectrometry through spatially targeted liquid micro-extractions
- PMID: 29226434
- PMCID: PMC5812809
- DOI: 10.1002/rcm.8042
Protein identification in imaging mass spectrometry through spatially targeted liquid micro-extractions
Abstract
Rationale: Liquid extraction surface analysis (LESA) can be used to generate spatially directed protein identifications in an imaging mass spectrometry (IMS) workflow. This approach involves the use of robotic micro-extractions coupled to online liquid chromatography (LC). We have characterized the extraction efficiency of this method as well as its ability to identify proteins from a matrix assisted laser/desorption ionization (MALDI) IMS experiment.
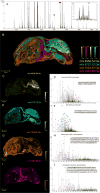
Methods: Proteins and peptides were extracted from transverse sections of a rat brain and sagittal sections of a mouse pup using liquid surface extractions. Extracts were either analyzed by online LC coupled to a high mass resolution Fourier transform ion cyclotron resonance (FTICR) mass spectrometer or collected offline and analyzed by traditional LC/MS methods. Identifications were made using both top-down and bottom-up methodologies. MALDI images were acquired on a 15T FTICR mass spectrometer at 125 μm spatial resolution.
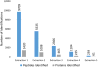
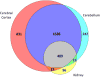
Results: Robotic liquid surface extractions are reproducible across various tissue types, providing significantly improved spatial resolution, with respect to extractions, while still allowing for a robust number of protein identifications. A single 2-μL extract can provide identification of over 14,000 peptides with little sample preparation, increasing throughput for spatially targeted workflows. Surface extractions from tissue were coupled directly to LC to gather spatially relevant proteomics data.
Conclusions: Robotic liquid surface extractions can be used to interrogate discrete regions of tissue to provide protein identifications with high throughput, accuracy, and robustness. The direct coupling of tissue surface extractions and LC offers a new and effective approach to provide spatial proteomics data in an imaging experiment.
Copyright © 2017 John Wiley & Sons, Ltd.
Figures
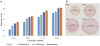
References
-
- Caprioli RM, Farmer TB, Gile J. Molecular imaging of biological samples: Localization of peptides and proteins using MALDI-TOF MS. Analytical Chemistry. 1997;69(23):4751–4760. - PubMed
-
- Meding S, Nitsche U, Balluff B, Elsner M, Rauser S, Schone C, Nipp M, Maak M, Feith M, Ebert MP, Friess H, Langer R, Hofler H, Zitzelsberger H, Rosenberg R, Walch A. Tumor Classification of Six Common Cancer Types Based on Proteomic Profiling by MALDI Imaging. Journal of Proteome Research. 2012;11(3):1996–2003. - PubMed
-
- Castellino S, Groseclose MR, Wagner D. MALDI imaging mass spectrometry: bridging biology and chemistry in drug development. Bioanalysis. 2011;3(21):2427–2441. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
