EGFR regulates macrophage activation and function in bacterial infection
- PMID: 27482886
- PMCID: PMC5004944
- DOI: 10.1172/JCI83585
EGFR regulates macrophage activation and function in bacterial infection
Abstract
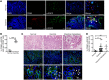
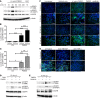
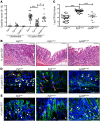
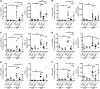
EGFR signaling regulates macrophage function, but its role in bacterial infection has not been investigated. Here, we assessed the role of macrophage EGFR signaling during infection with Helicobacter pylori, a bacterial pathogen that causes persistent inflammation and gastric cancer. EGFR was phosphorylated in murine and human macrophages during H. pylori infection. In human gastric tissues, elevated levels of phosphorylated EGFR were observed throughout the histologic cascade from gastritis to carcinoma. Deleting Egfr in myeloid cells attenuated gastritis and increased H. pylori burden in infected mice. EGFR deficiency also led to a global defect in macrophage activation that was associated with decreased cytokine, chemokine, and NO production. We observed similar alterations in macrophage activation and disease phenotype in the Citrobacter rodentium model of murine infectious colitis. Mechanistically, EGFR signaling activated NF-κB and MAPK1/3 pathways to induce cytokine production and macrophage activation. Although deletion of Egfr had no effect on DC function, EGFR-deficient macrophages displayed impaired Th1 and Th17 adaptive immune responses to H. pylori, which contributed to decreased chronic inflammation in infected mice. Together, these results indicate that EGFR signaling is central to macrophage function in response to enteric bacterial pathogens and is a potential therapeutic target for infection-induced inflammation and associated carcinogenesis.
Figures

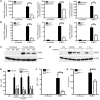



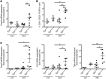
References
MeSH terms
Substances
Grants and funding
- R01 DK053620/DK/NIDDK NIH HHS/United States
- P01 CA028842/CA/NCI NIH HHS/United States
- P50 CA095103/CA/NCI NIH HHS/United States
- R01 CA077955/CA/NCI NIH HHS/United States
- R01 CA190612/CA/NCI NIH HHS/United States
- P30 DK058404/DK/NIDDK NIH HHS/United States
- R01 AT004821/AT/NCCIH NIH HHS/United States
- P01 CA116087/CA/NCI NIH HHS/United States
- I01 BX001453/BX/BLRD VA/United States
- P30 CA068485/CA/NCI NIH HHS/United States
- T32 GM008554/GM/NIGMS NIH HHS/United States
- R01 DK058587/DK/NIDDK NIH HHS/United States
- F31 DK107159/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

