Abstract
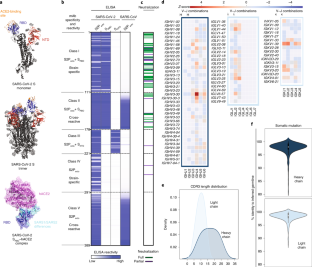
Antibodies are a principal determinant of immunity for most RNA viruses and have promise to reduce infection or disease during major epidemics. The novel coronavirus SARS-CoV-2 has caused a global pandemic with millions of infections and hundreds of thousands of deaths to date1,2. In response, we used a rapid antibody discovery platform to isolate hundreds of human monoclonal antibodies (mAbs) against the SARS-CoV-2 spike (S) protein. We stratify these mAbs into five major classes on the basis of their reactivity to subdomains of S protein as well as their cross-reactivity to SARS-CoV. Many of these mAbs inhibit infection of authentic SARS-CoV-2 virus, with most neutralizing mAbs recognizing the receptor-binding domain (RBD) of S. This work defines sites of vulnerability on SARS-CoV-2 S and demonstrates the speed and robustness of advanced antibody discovery platforms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and Supplementary Information. The ImMunoGeneTics database is available from http://www.imgt.org/. The analysis pipeline PyIR (https://github.com/crowelab/PyIR) and the specific scripts used for sequence analysis (https://github.com/crowelab/cov2-panel-scripts) are available. Structures deposited by other groups for the full-length spike trimer (6VYB) and the RBD–hACE2 complex (6M0J) that were used for visualization in this paper are publicly available (www.rcsb.org). Sequences for mAbs described in this study have been deposited at GenBank and are available under the following accession codes: MT665032–MT665070, MT665419–MT665457, MT665071–MT665418 and MT665458–MT665805. Datasets are available from the corresponding authors upon reasonable request.
References
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 382, 727–733 (2020).
Sui, J. et al. Potent neutralization of severe acute respiratory syndrome (SARS) coronavirus by a human mAb to S1 protein that blocks receptor association. Proc. Natl Acad. Sci. USA 101, 2536–2541 (2004).
ter Meulen, J. et al. Human monoclonal antibody as prophylaxis for SARS coronavirus infection in ferrets. Lancet 363, 2139–2141 (2004).
ter Meulen, J. et al. Human monoclonal antibody combination against SARS coronavirus: synergy and coverage of escape mutants. PLoS Med. 3, e237 (2006).
Zhu, Z. et al. Potent cross-reactive neutralization of SARS coronavirus isolates by human monoclonal antibodies. Proc. Natl Acad. Sci. USA 104, 12123–12128 (2007).
Rockx, B. et al. Structural basis for potent cross-neutralizing human monoclonal antibody protection against lethal human and zoonotic severe acute respiratory syndrome coronavirus challenge. J. Virol. 82, 3220–3235 (2008).
Chen, Z. et al. Human neutralizing monoclonal antibody inhibition of Middle East respiratory syndrome coronavirus replication in the common marmoset. J. Infect. Dis. 215, 1807–1815 (2017).
Choi, J. H. et al. Characterization of a human monoclonal antibody generated from a B-cell specific for a prefusion-stabilized spike protein of Middle East respiratory syndrome coronavirus. PLoS ONE 15, e0232757 (2020).
Niu, P. et al. Ultrapotent human neutralizing antibody repertoires against Middle East respiratory syndrome coronavirus from a recovered patient. J. Infect. Dis. 218, 1249–1260 (2018).
Wang, L., et al. Importance of neutralizing monoclonal antibodies targeting multiple antigenic sites on the Middle East respiratory syndrome coronavirus spike glycoprotein to avoid neutralization escape. J. Virol. 92, e02002-17 (2018).
Wang, N. et al. Structural definition of a neutralization-sensitive epitope on the MERS-CoV S1-NTD. Cell Rep. 28, e3396 (2019).
Zhang, S. et al. Structural definition of a unique neutralization epitope on the receptor-binding domain of MERS-CoV spike glycoprotein. Cell Rep. 24, 441–452 (2018).
Corti, D. et al. Prophylactic and postexposure efficacy of a potent human monoclonal antibody against MERS coronavirus. Proc. Natl Acad. Sci. USA 112, 10473–10478 (2015).
Jiang, L. et al. Potent neutralization of MERS-CoV by human neutralizing monoclonal antibodies to the viral spike glycoprotein. Sci. Transl. Med. 6, 234ra259 (2014).
Tang, X. C. et al. Identification of human neutralizing antibodies against MERS-CoV and their role in virus adaptive evolution. Proc. Natl Acad. Sci. USA 111, E2018–E2026 (2014).
Ying, T. et al. Exceptionally potent neutralization of Middle East respiratory syndrome coronavirus by human monoclonal antibodies. J. Virol. 88, 7796–7805 (2014).
Jiang, S., Hillyer, C. & Du, L. Neutralizing antibodies against SARS-CoV-2 and other human coronaviruses. Trends Immunol. 41, 355–359 (2020).
Gilchuk, P. et al. Integrated technology platform for accelerated discovery of antiviral antibody therapeutics. Nat. Biomed. Eng. (in the press).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Holshue, M. L. et al. First case of 2019 novel coronavirus in the United States. N. Engl. J. Med. 382, 929–936 (2020).
Gilchuk, P. et al. Analysis of a therapeutic antibody cocktail reveals determinants for cooperative and broad ebolavirus meutralization. Immunity 52, e312 (2020).
Soto, C. et al. High frequency of shared clonotypes in human B cell receptor repertoires. Nature 566, 398–402 (2019).
Wrammert, J. et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J. Exp. Med. 208, 181–193 (2011).
Case, J. B. et al. Neutralizing antibody and soluble ACE2 inhibition of a replication-competent VSV-SARS-CoV-2 and a clinical isolate of SARS-CoV-2. Cell Host Microbe https://doi.org/10.1016/j.chom.2020.06.021 (2020).
Rogers, T. F. et al. Rapid isolation of potent SARS-CoV-2 neutralizing antibodies and protection in a small animal model. Science https://doi.org/10.1126/science.abc7520 (2020).
Robbiani, D. F. et al. Convergent antibody responses to SARS-CoV-2 infection in convalescent individuals. Nature https://doi.org/10.1038/s41586-020-2456-9 (2020).
Brouwer, P. J. M. et al. Potent neutralizing antibodies from COVID-19 patients define multiple targets of vulnerability. Science eabc5902 (2020).
Cao, Y. et al. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent patients’ B cells. Cell https://doi.org/10.1016/j.cell.2020.05.025 (2020).
Shi, R. et al. A human neutralizing antibody targets the receptor binding site of SARS-CoV-2. Nature https://doi.org/10.1038/s41586-020-2381-y (2020).
Wu, Y. et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 368, 1274–1278 (2020).
Ju, B. et al. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature https://doi.org/10.1038/s41586-020-2380-z (2020).
Wec, A. Z. et al. Broad neutralization of SARS-related viruses by human monoclonal antibodies. Science https://doi.org/10.1126/science.abc7424 (2020).
Williamson, L. E. et al. Early human B cell response to Ebola virus in four U.S. survivors of infection. J. Virol. 93, e01439-18 (2019).
Davis, C. W. et al. Longitudinal analysis of the human B cell response to Ebola virus infection. Cell 177, e1517 (2019).
Zost, S. J. et al. Potently neutralizing human antibodies that block SARS-CoV-2 receptor binding and protect animals. Nature (in the press).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, e286 (2020).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Mukherjee, S. et al. Enhancing dengue virus maturation using a stable furin over-expressing cell line. Virology 497, 33–40 (2016).
Ohi, M., Li, Y., Cheng, Y. & Walz, T. Negative staining and image classification—powerful tools in modern electron microscopy. Biol. Proced. Online 6, 23–34 (2004).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T., Noble, A. J. & Berger, B. Topaz-Denoise: general deep denoising models for cryoEM. Preprint at https://doi.org/10.1101/838920 (2019).
Nguyen, D. C. et al. Factors of the bone marrow microniche that support human plasma cell survival and immunoglobulin secretion. Nat. Commun. 9, 3698 (2018).
Guthmiller, J. J., Dugan, H. L., Neu, K. E., Lan, L. Y. & Wilson, P. C. An efficient method to generate monoclonal antibodies from human B cells. Methods Mol. Biol. 1904, 109–145 (2019).
Soto C. F. J. et al. PyIR: A scalable wrapper for processing billions of immunoglobulin and T cell receptor sequences using IgBLAST. BMC Bioinformatics (in the press).
Chng, J. et al. Cleavage efficient 2A peptides for high level monoclonal antibody expression in CHO cells. MAbs 7, 403–412 (2015).
Scobey, T. et al. Reverse genetics with a full-length infectious cDNA of the Middle East respiratory syndrome coronavirus. Proc. Natl Acad. Sci. USA 110, 16157–16162 (2013).
Yount, B. et al. Reverse genetics with a full-length infectious cDNA of severe acute respiratory syndrome coronavirus. Proc. Natl Acad. Sci. USA 100, 12995–13000 (2003).
Acknowledgements
We thank M. Mayo and N. S. Galeano for coordination of human studies, D. O’Connor, N. Safdar, G. Baird, J. Shendure and S. Mubareka for helpful advice on human patients, A. Jones and the staff of the Vanderbilt VANTAGE core laboratory for expedited sequencing, R. Trosseth for assistance with data management and analysis, R. Bombardi and C. Soto for technical consultation on genomics approaches, A. Kim for production of a recombinant form of the mAb CR3022, C. Swearingen and the staff of Fedex Express Specialty Services for expedited transport services, V. Pai and K. Breinlinger of Berkeley Lights, and K. Louder and scientists at Twist Bioscience, B. Fritz at 10x Genomics and representatives at ACEA Biosciences for providing resources, outstanding expedited services and expert applications support. We thank A. Ward, S. Bangaru, P. McTamney, K. Ren and A. Barnes for protein reagents. This study was supported by Defense Advanced Research Projects Agency (DARPA) grant HR0011-18-2-0001, NIH contracts 75N93019C00074 and 75N93019C00062 and the Dolly Parton COVID-19 Research Fund at Vanderbilt. This work was supported by NIH grant 1S10RR028106-01A1 for the Next Generation Nucleic Acid Sequencer, housed in Vanderbilt Technologies for Advanced Genomics (VANTAGE) and supported by the National Center for Research Resources, grant UL1 RR024975-01, and is now at the National Center for Advancing Translational Sciences, grant 2 UL1 TR000445-06. S.J.Z. was supported by NIH T32 AI095202. J.B.C. is supported by a Helen Hay Whitney Foundation postdoctoral fellowship. D.R.M. was supported by NIH T32 AI007151 and a Burroughs Wellcome Fund Postdoctoral Enrichment Program Award. J.E.C. is the recipient of the 2019 Future Insight Prize from Merck KGaA, which supported this research with a research grant. The content is solely the responsibility of the authors and does not necessarily represent the official views of the US government or the other sponsors.
Author information
Authors and Affiliations
Contributions
Conceived of the project: S.J.Z., P.G., R.H.C., L.B.T., M.S.D., J.E.C.; Obtained funding: J.E.C. and M.S.D. Obtained human samples: M.O., H.Y.C., J.E.C.; Performed laboratory experiments: S.J.Z., P.G., R.E.C., J.B.C., J.X.R., A.T., R.S.N., R.E.S., N.S., E.B., J.E.D., K.W.M., S.S., D.R.M, P.W.R., L.-M.B.; Performed computational work: E.C.C., T.J., S.D., L.M.; Supervised research: S.P.J.W. M.S.D., L.B.T., R.S.B., R.H.C., J.E.C.; Provided critical reagents: J.E.D., K.W.M., F.E.-H.L., D.C.N., I.S., R.S.B.; Wrote the first draft of the paper: S.J.Z., P.G., R.H.C., J.E.C. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.S.B. has served as a consultant for Takeda and Sanofi Pasteur on issues related to vaccines. M.S.D. is a consultant for Inbios, Vir Biotechnology, NGM Biopharmaceuticals and Eli Lilly, is on the scientific advisory board of Moderna and is a recipient of unrelated research grants from Moderna and Emergent BioSolutions. H.Y.C. has served as a consultant for Merck and GlaxoSmithKline and has received research funding from Sanofi Pasteur and research support from Cepheid, Genentech and Ellume. J.E.C. has served as a consultant for Sanofi and is on the scientific advisory boards of CompuVax and Meissa Vaccines, is a recipient of previous unrelated research grants from Moderna and Sanofi and is founder of IDBiologics. Vanderbilt University has applied for patents concerning SARS-CoV-2 antibodies that are related to this work. Emory University has applied for a patent concerning the plasmablast survival medium. S.P.J.W. and P.W.R. have filed a disclosure with Washington University for the recombinant VSV. J.E.D. and K.W.M. are employees of Berkeley Lights. All other authors declared no competing interests.
Additional information
Peer review information Joao Monteiro was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Expression and validation of prefusion-stabilized SARS-CoV-2 S2Pecto protein.
a. Reducing SDS-PAGE gel indicating S2Pecto protein migrating at approximately 180 KDa. One representative protein preparation is shown. b. Representative micrograph of negative-stain electron microscopy with S2Pecto protein preparation. Scale bar denotes 100 nm. c. 2D class-averages of S2Pecto protein in the prefusion conformation. The size for each box is 128 pixels. Detailed information on image collection is available in Supplementary Table 2.
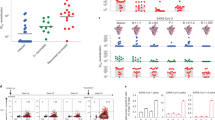
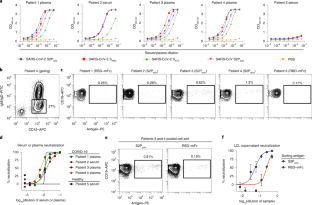
Extended Data Fig. 2 Representative gating strategy for antigen-specific cell sorting.
Representative gating strategy for profiling of antigen-specific B cell frequency for donors. Subject 4 is shown, and phenotypic markers are shown on plot axes. Arrows indicate cell populations derived from gates.
Extended Data Fig. 3 Functional assays from single antigen-reactive B cells.
a. Biotinylated antigen (dark grey) was coupled to a streptavidin-conjugated polystyrene bead (light grey). Antibodies (blue) are secreted by single B cells loaded into individual NanoPens on the Berkeley Lights Beacon optofluidic device. Antibody binding to antigen was detected with a fluorescent anti-human IgG secondary Ab (black). b. Left: Schematic of fluorescing beads in the channel above a pen containing an individual B cell indicates antigen-specific reactivity. Top right: False-color still image of positive wells with B cells secreting S2Pecto-reactive antibodies. Reactive antibody diffusing out of a pen is visualized as a plume of fluorescence. Bottom right: False-color still image of positive wells with B cells secreting RBD-mFc-reactive antibodies. c. Representative images of RBD-mFc reactive B cells from a single-B-cell secretion assay. d. Identification of mAbs with hACE2-blocking activity using single-cell functional screening. Left: Schematic illustrating detection of secreted Ab and hACE2 binding on an RBD-mFc-coated streptavidin bead. Ab binding was detected in one fluorescent channel, while hACE2 binding was detected in another fluorescent channel. The top panel illustrates an RBD-binding, non-blocking mAb, where the bead is positive for both Ab and hACE2 signals, while the bottom panel illustrates an RBD-binding mAb that competes with hACE2 for binding, where the bead is positive for only Ab signal. Right: Representative images of a B cell secreting non-blocking Abs (top) and a B cell secreting hACE2-blocking mAbs (bottom). Streptavidin beads are loaded into the same pens as B cells. The fluorescence of the streptavidin beads in the same pen as the B cell secreting hACE2-blocking Abs is reduced relative to adjacent wells, indicating hACE2-blocking activity.
Extended Data Fig. 4 Real-time cell analysis assay to screen for neutralization activity.
a. Curves for fully neutralizing mAb (green) and partially neutralizing mAb (red) by monitoring of CPE in Vero-furin cells that were inoculated with SARS-CoV-2 and pre-incubated with the respective mAb. Uninfected cells (blue) and infected cells without antibody addition (grey) served as controls for intact monolayer and full CPE, respectively. Data represent a single well measurement for each mAb at the highest tested concentration, mean ± SD values of technical duplicates for the positive CPE control, and mean ± SD values of technical quadruplicates for the no-CPE controls. b. Example sensograms from individual wells of 384-well E-plate analysis showing rapid identification of SARS-CoV-2 neutralizing mAbs. Neutralization was assessed using micro-scale purified mAbs and each mAb was tested in four 5-fold dilutions as indicated. Plates were measured every 8–12 hrs for a total of 72 hrs as in (a).
Extended Data Fig. 5 Real-time cell analysis assay to quantify neutralization potency.
Dose-response curves showing activity of neutralizing mAbs that were identified by rapid screening using the RTCA assay, as in Extended Data Fig. 4. Each mAb was tested in four sequential five-fold dilutions from micro-scale purified samples in which mAbs concentrations were not normalized but quantified. Neutralization was calculated as the percent of maximal cell index in control wells without virus minus cell index in control (virus-only) wells that exhibited maximal CPE at 40 to 48 hrs after applying virus-antibody mixture to the cells. a. Representative neutralizing mAbs that fully prevented CPE at the lowest tested dilution (corresponding to the highest tested mAb concentration) are shown. IC50 values estimated from each curve are indicated. Curves for potently neutralizing mAbs (IC50 < 100 ng/mL) are shown in orange, from which mAbs COV2–2355 and COV2–2381 are genetically related. b. Representative neutralizing mAbs that partially prevented CPE at the lowest tested dilution (corresponding to the highest tested mAb concentration) are shown.
Extended Data Fig. 6 Quantitative neutralization assays of VSV-SARS-CoV-2.
Dose-response neutralization of VSV-SARS-CoV-2 by neutralizing mAbs. IC50 values are indicated for each mAb. Data shown are the mean of two technical replicates from a single experiment, and error bars denote the standard deviation for each point.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
41591_2020_998_MOESM3_ESM.mov
Supplementary Video 1 Time-lapse imaging of antigen-sorted single B cells secreting S2Pecto-reactive antibodies. A field of view of the optofluidic chip is shown with single B cells at the bottom of NanoPens, as in Extended Data Fig. 3. Antigen-reactive antibody bound to S2Pecto antigen conjugated to streptavidin polystyrene beads loaded into the channel is detected by an anti-IgG secondary antibody. Positive wells are identified by the specific bloom of fluorescence signal, indicating antigen-specific antibody diffusing out of a single pen. The edges of pens are highlighted in green, and pen numbers are shown in yellow. For that field of view, there were 96 pens containing IgG-secreting B cells, with 53 cells secreting trimer-reactive antibody and 30 cells secreting antibody reactive to RBD, giving an overall IgG antigen-reactive frequency of approximately 55%. The movie is composed of still images obtained every 5 min over the course of a 30-min assay.
Supplementary Table 4
SARS-CoV-2 mAb reactivity, neutralization and sequence features.
Rights and permissions
About this article
Cite this article
Zost, S.J., Gilchuk, P., Chen, R.E. et al. Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein. Nat Med 26, 1422–1427 (2020). https://doi.org/10.1038/s41591-020-0998-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-020-0998-x
This article is cited by
-
High-throughput strategies for monoclonal antibody screening: advances and challenges
Journal of Biological Engineering (2025)
-
NIEAs elicited by wild-type SARS-CoV-2 primary infection fail to enhance the infectivity of Omicron variants
Virology Journal (2025)
-
Enhancing large particle recovery in high-throughput functional cell sorting through ΔBOP optimization
Scientific Reports (2025)
-
A novel anti-HER2/EGFR bispecific antibody–drug conjugate demonstrates promising antitumor efficacy and overcomes resistance to HER2- or EGFR-targeted ADCs
Investigational New Drugs (2025)
-
Rapid and highly sensitive immunoassay using an ultra-thin immuno-wall microfluidic device with a sequential fluorescence signal increment method
Analytical and Bioanalytical Chemistry (2025)



