Specificity and affinity of the N-terminal residues in staphylocoagulase in binding to prothrombin
- PMID: 32156702
- PMCID: PMC7186164
- DOI: 10.1074/jbc.RA120.012588
Specificity and affinity of the N-terminal residues in staphylocoagulase in binding to prothrombin
Abstract
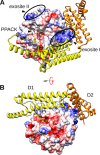
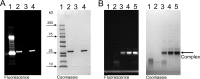
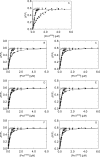
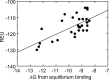
In Staphylococcus aureus-caused endocarditis, the pathogen secretes staphylocoagulase (SC), thereby activating human prothrombin (ProT) and evading immune clearance. A previous structural comparison of the SC(1-325) fragment bound to thrombin and its inactive precursor prethrombin 2 has indicated that SC activates ProT by inserting its N-terminal dipeptide Ile1-Val2 into the ProT Ile16 pocket, forming a salt bridge with ProT's Asp194, thereby stabilizing the active conformation. We hypothesized that these N-terminal SC residues modulate ProT binding and activation. Here, we generated labeled SC(1-246) as a probe for competitively defining the affinities of N-terminal SC(1-246) variants preselected by modeling. Using ProT(R155Q,R271Q,R284Q) (ProTQQQ), a variant refractory to prothrombinase- or thrombin-mediated cleavage, we observed variant affinities between ∼1 and 650 nm and activation potencies ranging from 1.8-fold that of WT SC(1-246) to complete loss of function. Substrate binding to ProTQQQ caused allosteric tightening of the affinity of most SC(1-246) variants, consistent with zymogen activation through occupation of the specificity pocket. Conservative changes at positions 1 and 2 were well-tolerated, with Val1-Val2, Ile1-Ala2, and Leu1-Val2 variants exhibiting ProTQQQ affinity and activation potency comparable with WT SC(1-246). Weaker binding variants typically had reduced activation rates, although at near-saturating ProTQQQ levels, several variants exhibited limiting rates similar to or higher than that of WT SC(1-246). The Ile16 pocket in ProTQQQ appears to favor nonpolar, nonaromatic residues at SC positions 1 and 2. Our results suggest that SC variants other than WT Ile1-Val2-Thr3 might emerge with similar ProT-activating efficiency.
Keywords: Staphylococcus aureus (S. aureus); affinity; clot formation; coagulation; coagulation factor; competitive equilibrium binding; endocarditis; equilibrium binding; fibrin; kinetics; ligand-binding protein; prothrombin; specificity; staphylocoagulase (SC); virulence factor.
© 2020 Maddur et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures





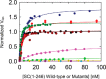

References
-
- Hartleib J., Köhler N., Dickinson R. B., Chhatwal G. S., Sixma J. J., Hartford O. M., Foster T. J., Peters G., Kehrel B. E., and Herrmann M. (2000) Protein A is the von Willebrand factor binding protein on Staphylococcus aureus. Blood 96, 2149–2156 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

