Single Cell Profiling in the Sox10Dom Hirschsprung Mouse Implicates Hox Genes in Enteric Neuron Trajectory Allocation
- PMID: 40701368
- PMCID: PMC12494863
- DOI: 10.1016/j.jcmgh.2025.101590
Single Cell Profiling in the Sox10Dom Hirschsprung Mouse Implicates Hox Genes in Enteric Neuron Trajectory Allocation
Abstract
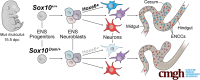
Background & aims: Enteric nervous system (ENS) development requires migration, proliferation, and differentiation of progenitors for normal gastrointestinal (GI) motility. Sox10 deficit causes aganglionosis, modeling Hirschsprung disease (HSCR), and disrupts ratios of postnatal enteric neurons in proximal ganglionated bowel. How Sox10 deficiency alters enteric neuron ratios is unclear. Sox10's prominent expression in enteric neural crest-derived progenitors (ENCPs) and lack of this gene in mature enteric neurons led us to examine Sox10Dom effects in early ENS development.
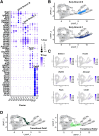
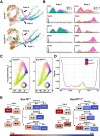
Methods: Immunohistochemistry localized SOX10 in the developing ENS relative to HuC/D. ENS progenitors, developing neurons, and enteric glia were isolated from Sox10+/+ and Sox10Dom/+ littermates for single-cell RNA sequencing (scRNA-seq). scRNA-seq data was processed to identify cell type-specific markers, differentially expressed genes, cell fate trajectories, and gene regulatory network activity between genotypes. Hybridization chain reaction (HCR) coupled with immunohistochemistry validated expression changes.
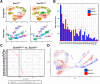
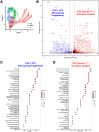
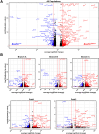
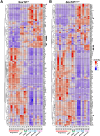
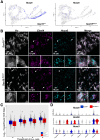
Results: SOX10 protein was detected in early ENS neurons. scRNA-seq profiles detected three neuronal trajectories emerging via two transition pathways accompanied by elevated activity of Hox gene regulatory networks (GRN). Sox10Dom/+ scRNA-seq profiles exhibited a novel progenitor cluster, reduced numbers of cells in transitional states, and shifts in cell abundance between neuronal trajectories. Hoxa6 was differentially expressed in the neuronal trajectories impacted in Sox10Dom/+ mutants, and HCR identified altered Hoxa6 expression in early developing neurons of Sox10Dom/+ ENS.
Conclusions: Sox10Dom/+ mutation shifts enteric neuron types by altering neuronal trajectories early in ENS development. Multiple neurogenic transcription factors are reduced in Sox10Dom/+ scRNA-seq profiles. This work is the first to correlate changes in Hox expression, notably Hoxa6, with alterations in enteric neuron trajectories.
Keywords: Enteric Nervous System; Hirschsprung Disease; Hoxa6; Neural Crest; Sox10.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Figures






Update of
-
Single Cell Profiling in the Sox10 Dom/+ Hirschsprung Mouse Implicates Hoxa6 in Enteric Neuron Lineage Allocation.bioRxiv [Preprint]. 2024 Sep 19:2024.09.18.613729. doi: 10.1101/2024.09.18.613729. bioRxiv. 2024. Update in: Cell Mol Gastroenterol Hepatol. 2025;19(12):101590. doi: 10.1016/j.jcmgh.2025.101590. PMID: 39345473 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

