Thymic epithelial cells require lipid kinase Vps34 for CD4 but not CD8 T cell selection
- PMID: 35997680
- PMCID: PMC9402993
- DOI: 10.1084/jem.20212554
Thymic epithelial cells require lipid kinase Vps34 for CD4 but not CD8 T cell selection
Abstract
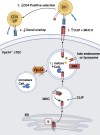
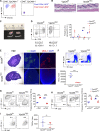
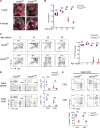
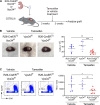
The generation of a functional, self-tolerant T cell receptor (TCR) repertoire depends on interactions between developing thymocytes and antigen-presenting thymic epithelial cells (TECs). Cortical TECs (cTECs) rely on unique antigen-processing machinery to generate self-peptides specialized for T cell positive selection. In our current study, we focus on the lipid kinase Vps34, which has been implicated in autophagy and endocytic vesicle trafficking. We show that loss of Vps34 in TECs causes profound defects in the positive selection of the CD4 T cell lineage but not the CD8 T cell lineage. Utilizing TCR sequencing, we show that T cell selection in conditional mutants causes altered repertoire properties including reduced clonal sharing. cTECs from mutant mice display an increased abundance of invariant chain intermediates bound to surface MHC class II molecules, indicating altered antigen processing. Collectively, these studies identify lipid kinase Vps34 as an important contributor to the repertoire of selecting ligands processed and presented by TECs to developing CD4 T cells.
© 2022 Postoak et al.
Conflict of interest statement
Disclosures: L. Van Kaer reported “I am a member of the scientific advisory board of Isu Abxis Co., Ltd. (South Korea), for which I get financial compensation. The company focuses on the development of antibody-mediated treatment for cancer and rare diseases. The work at the company is not directly related to this publication.” No other disclosures were reported.
Figures





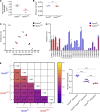
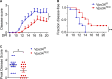
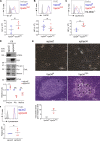
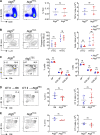
References
-
- Akiyama, T., Shimo Y., Yanai H., Qin J., Ohshima D., Maruyama Y., Asaumi Y., Kitazawa J., Takayanagi H., Penninger J.M., et al. 2008. The tumor necrosis factor family receptors RANK and CD40 cooperatively establish the thymic medullary microenvironment and self-tolerance. Immunity. 29:423–437. 10.1016/j.immuni.2008.06.015 - DOI - PubMed
-
- Barthlott, T., Kohler H., and Eichmann K.. 1997. Asynchronous coreceptor downregulation after positive thymic selection: Prolonged maintenance of the double positive state in CD8 lineage differentiation due to sustained biosynthesis of the CD4 coreceptor. J. Exp. Med. 185:357–362. 10.1084/jem.185.2.357 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- I01 BX001444/BX/BLRD VA/United States
- IK6 BX004595/BX/BLRD VA/United States
- P30 DK058404/DK/NIDDK NIH HHS/United States
- R01 AI139046/AI/NIAID NIH HHS/United States
- P30 CA068485/CA/NCI NIH HHS/United States
- R01 DK104817/DK/NIDDK NIH HHS/United States
- R01 AI137082/AI/NIAID NIH HHS/United States
- I01 BX006010/BX/BLRD VA/United States
- P30 DK020593/DK/NIDDK NIH HHS/United States
- 19TPA34910078/AHA/American Heart Association-American Stroke Association/United States
- T32 AR059039/AR/NIAMS NIH HHS/United States
- T32 HL069765/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

