Altered Osteoblast Metabolism with Aging Results in Lipid Accumulation and Oxidative Stress Mediated Bone Loss
- PMID: 37548937
- PMCID: PMC10917552
- DOI: 10.14336/AD.2023.0510
Altered Osteoblast Metabolism with Aging Results in Lipid Accumulation and Oxidative Stress Mediated Bone Loss
Abstract
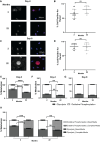
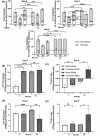
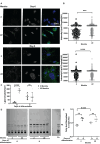
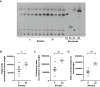
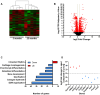
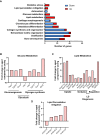
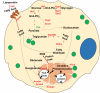
Cellular aging is associated with dysfunction of numerous tissues affecting multiple organ systems. A striking example of this is related to age-related bone loss, or osteoporosis, increasing fracture incidence. Interestingly, the two compartments of bone, cortical and cancellous or trabecular, rely on different mechanisms for development and maintenance during 'normal' aging. At a cellular level, the aging process disturbs a multitude of intracellular pathways. In particular, alterations in cellular metabolic functions thereby impacting cellular bioenergetics have been implicated in multiple tissues. Therefore, this study aimed to characterize how metabolic processes were altered in bone forming osteoblasts in aged mice compared to young mice. Metabolic flux analyses demonstrated both stromal cells and mature, matrix secreting osteoblasts from aged mice exhibited mitochondrial dysfunction. This was also accompanied by a lack of adaptability or metabolic flexibility to utilize exogenous substrates compared to osteoblasts cultured from young mice. Additionally, lipid droplets accumulated in both early stromal cells and mature osteoblasts from aged mice, which was further depicted as increased lipid content within the bone cortex of aged mice. Global transcriptomic analysis of the bone further supported these metabolic data as enhanced oxidative stress genes were up-regulated in aged mice, while osteoblast-related genes were down-regulated when compared to the young mice. Collectively, these data suggest that aging results in altered osteoblast metabolic handling of both exogenous and endogenous substrates which could contribute to age-related osteoporosis.
Conflict of interest statement
Authors have nothing to declare that could be constructed as potential conflict of interest.
Figures

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
