A Quantitative Framework for Evaluating Single-Cell Data Structure Preservation by Dimensionality Reduction Techniques
- PMID: 32375029
- PMCID: PMC7305633
- DOI: 10.1016/j.celrep.2020.107576
A Quantitative Framework for Evaluating Single-Cell Data Structure Preservation by Dimensionality Reduction Techniques
Abstract
High-dimensional data, such as those generated by single-cell RNA sequencing (scRNA-seq), present challenges in interpretation and visualization. Numerical and computational methods for dimensionality reduction allow for low-dimensional representation of genome-scale expression data for downstream clustering, trajectory reconstruction, and biological interpretation. However, a comprehensive and quantitative evaluation of the performance of these techniques has not been established. We present an unbiased framework that defines metrics of global and local structure preservation in dimensionality reduction transformations. Using discrete and continuous real-world and synthetic scRNA-seq datasets, we show how input cell distribution and method parameters are largely determinant of global, local, and organizational data structure preservation by 11 common dimensionality reduction methods.
Keywords: data analysis; dimensionality reduction; single-cell analysis; single-cell transcriptomics; unsupervised learning; visualization.
Copyright © 2020 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures



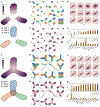
References
-
- Becht E, McInnes L, Healy J, Dutertre C-A, Kwok IWH, Ng LG, Ginhoux F, and Newell EW (2018). Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol 37, 38–44. - PubMed
-
- Cramér H (1928). On the composition of elementary errors. Scand. Actuar. J 1928, 13–74.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

