Improving lung cancer diagnosis with cancer, fungal, and imaging biomarkers
- PMID: 36792410
- PMCID: PMC10287834
- DOI: 10.1016/j.jtcvs.2022.12.014
Improving lung cancer diagnosis with cancer, fungal, and imaging biomarkers
Abstract
Objective: Indeterminate pulmonary nodules (IPNs) represent a significant diagnostic burden in health care. We aimed to compare a combination clinical prediction model (Mayo Clinic model), fungal (histoplasmosis serology), imaging (computed tomography [CT] radiomics), and cancer (high-sensitivity cytokeratin fraction 21; hsCYFRA 21-1) biomarker approach to a validated prediction model in diagnosing lung cancer.
Methods: A prospective specimen collection, retrospective blinded evaluation study was performed in 3 independent cohorts with 6- to 30-mm IPNs (n = 281). Serum histoplasmosis immunoglobulin G and immunoglobulin M antibodies and hsCYFRA 21-1 levels were measured and a validated CT radiomic score was calculated. Multivariable logistic regression models were estimated with Mayo Clinic model variables, histoplasmosis antibody levels, CT radiomic score, and hsCYFRA 21-1. Diagnostic performance of the combination model was compared with that of the Mayo Clinic model. Bias-corrected clinical net reclassification index (cNRI) was used to estimate the clinical utility of a combination biomarker approach.
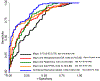
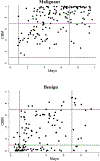
Results: A total of 281 patients were included (111 from a histoplasmosis-endemic region). The combination biomarker model including the Mayo Clinic model score, histoplasmosis antibody levels, radiomics, and hsCYFRA 21-1 level showed improved diagnostic accuracy for IPNs compared with the Mayo Clinic model alone with an area under the receiver operating characteristics curve of 0.80 (95% CI, 0.76-0.84) versus 0.72 (95% CI, 0.66-0.78). Use of this combination model correctly reclassified intermediate risk IPNs into low- or high-risk category (cNRI benign = 0.11 and cNRI malignant = 0.16).
Conclusions: The addition of cancer, fungal, and imaging biomarkers improves the diagnostic accuracy for IPNs. Integrating a combination biomarker approach into the diagnostic algorithm of IPNs might decrease unnecessary invasive testing of benign nodules and reduce time to diagnosis for cancer.
Keywords: biomarker; cancer; nodule; pulmonary.
Copyright © 2023. Published by Elsevier Inc.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

