Unsupervised Trajectory Analysis of Single-Cell RNA-Seq and Imaging Data Reveals Alternative Tuft Cell Origins in the Gut
- PMID: 29153838
- PMCID: PMC5799016
- DOI: 10.1016/j.cels.2017.10.012
Unsupervised Trajectory Analysis of Single-Cell RNA-Seq and Imaging Data Reveals Alternative Tuft Cell Origins in the Gut
Abstract
Modern single-cell technologies allow multiplexed sampling of cellular states within a tissue. However, computational tools that can infer developmental cell-state transitions reproducibly from such single-cell data are lacking. Here, we introduce p-Creode, an unsupervised algorithm that produces multi-branching graphs from single-cell data, compares graphs with differing topologies, and infers a statistically robust hierarchy of cell-state transitions that define developmental trajectories. We have applied p-Creode to mass cytometry, multiplex immunofluorescence, and single-cell RNA-seq data. As a test case, we validate cell-state-transition trajectories predicted by p-Creode for intestinal tuft cells, a rare, chemosensory cell type. We clarify that tuft cells are specified outside of the Atoh1-dependent secretory lineage in the small intestine. However, p-Creode also predicts, and we confirm, that tuft cells arise from an alternative, Atoh1-driven developmental program in the colon. These studies introduce p-Creode as a reliable method for analyzing large datasets that depict branching transition trajectories.
Keywords: cell-state transitions; differentiation hierachies; graph theory; intestine and colon; mass cytometry; pseudo-time analysis; single-cell RNA-seq; single-cell biology; trajectories; tuft cells.
Copyright © 2017 The Authors. Published by Elsevier Inc. All rights reserved.
Figures


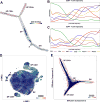
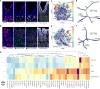

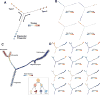

Comment in
-
Breaking New Ground in the Landscape of Single-Cell Analysis.Cell Syst. 2018 Jan 24;6(1):5-7. doi: 10.1016/j.cels.2017.12.015. Cell Syst. 2018. PMID: 29401450
References
-
- Anchang B, Hart TDP, Bendall SC, Qiu P, Bjornson Z, Linderman M, Nolan GP, Plevritis SK. Visualization and cellular hierarchy inference of single-cell data using SPADE. Nat. Protoc. 2016;11:1264–1279. - PubMed
-
- Barker N, van Es JH, Kuipers J, Kujala P, van den Born M, Cozijnsen M, Haegebarth A, Korving J, Begthel H, Peters PJ, et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature. 2007;449:1003–1007. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

