Differentially expressed microRNAs in the aqueous humor of patients with exfoliation glaucoma or primary open-angle glaucoma
- PMID: 29401312
- PMCID: PMC6048986
- DOI: 10.1093/hmg/ddy040
Differentially expressed microRNAs in the aqueous humor of patients with exfoliation glaucoma or primary open-angle glaucoma
Abstract
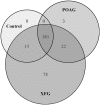
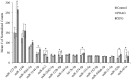
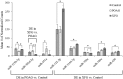
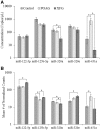
Both exfoliation glaucoma (XFG) and primary open-angle glaucoma (POAG) have been linked to decreased conventional outflow of aqueous humor (AH). To better understand the molecular changes in the AH content under such conditions, we analyzed the miRNA profiles of AH samples from patients with POAG and XFG compared to non-glaucoma controls. Individual AH samples (n = 76) were collected from POAG and XFG patients and age-matched controls during surgical procedure. After RNA extraction, the miRNA profiles were individually determined in 12 POAG, 12 XFG and 11 control samples. We identified 205, 295 and 195 miRNAs in the POAG, XFG and control samples, respectively. Our differential expression analysis identified three miRNAs (miR-125b-5p, miR-302d-3p and miR-451a) significantly different between POAG and controls, five miRNAs (miR-122-5p, miR-3144-3p, miR-320a, miR-320e and miR-630) between XFG and controls and one miRNA (miR-302d-3p) between POAG and XFG. While none of these miRNAs have been previously linked to glaucoma, miR-122-5p may target three glaucoma-associated genes: OPTN, TMCO1 and TGF-ß1. Pathway analysis revealed that these miRNAs are involved in potential glaucoma pathways, including focal adhesion, tight junctions, and TGF-ß signaling. Comparison of the miRNA profile in AH to unrelated human serum (n = 12) exposed potential relationships between these two fluids, although they were not significantly correlated. In summary, we have successfully profiled the miRNA expression without amplification in individual human AH samples and identified several POAG or XFG-associated miRNAs. These miRNAs may play a role in pathways previously implicated in glaucoma and act as biomarkers for disease pathogenesis.
Figures


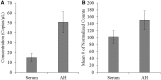
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

